St. Jude Children’s Research Hospital is known for going the distance in its mission of “Finding cures. Saving children.” This winter that included a 1,000-plus mile trek across five states to track and learn about Highly Pathogenic Avian Influenza in the Central Flyway.
Avian influenza viruses, including strains such as Highly Pathogenic Avian Influenza (HPAI), have not demonstrated sustained human-to-human transmission. The current public health risk associated with H5N1 avian influenza remains low.
The public health risk of avian influenza may be low, but patients receiving care at St. Jude are already fighting life-threatening diseases. There, any risk is already too high.
“St. Jude is focused on catastrophic diseases like cancer and rare genetic diseases,” said Lisa Kercher, director of The Webby Group lab operations for a St. Jude research team. “In the space of treating those diseases, the patient’s immune system may become compromised, leaving them vulnerable to adverse outcomes of infectious diseases like influenza."
To learn more about how bird flu viruses emerge and are transmitted, the St. Jude Center of Excellence for Influenza Research and Response (SJCEIRR) has been surveilling strains of avian influenza for decades so that medical teams have a leg up in an emergency situation.
“The job of St. Jude is to try to close some of the gaps in the research.”
Kercher’s work to safeguard medically vulnerable children — particularly through research on avian influenza and highly pathogenic strains such as H5N1 — has advanced understanding of zoonotic transmission dynamics and viral evolution at the human–wildlife interface. By integrating surveillance data from wild bird populations with clinical and epidemiological insights relevant to pediatric health, this work contributes to earlier detection of emerging strains, improved risk assessment, and more effective prevention strategies aimed at protecting immunocompromised and hospitalized children. Simultaneously, these efforts provide natural resource biologists with critical information on infection patterns, host ecology, and environmental drivers of disease persistence in wild avian populations.
"The current circulating strain of HPAI, H5N1, has circumvented the globe pretty rapidly and is making more wild birds sick than ever," Kercher said.
To better test for and analyze emerging influenza viruses, Kercher has developed a mobile diagnostic lab, known as BIRDWATCH, that allows for on-site evaluation and genetic sequencing of bird samples and provides real-time data to her team as well as wildlife managers. Prior to this mobile strategy for sample collection and analysis, it could take months for the results to filter back to the wildlife managers, during which time the virus may have already changed.
Reassortment: How Viruses Change
“If two different influenza viruses infect the same bird, they can swap genetic material and produce an entirely new virus,” said Kercher. “That new strain might be less harmful, just as severe, or potentially more dangerous than the original viruses — and we still can’t reliably predict which it will be.”
Avian influenza viruses contain eight distinct RNA gene segments that function like interchangeable building blocks. Different strains carry different combinations of these segments, giving each one a unique genetic profile. When two strains infect a single host, these segments can reshuffle — a process known as genetic reassortment — resulting in a newly mixed virus with its own characteristics and potential risks.
"The hope is to help biologists make better risk management decisions," Kercher said. "There are unique risk management challenges in each state depending on the geography and what kind of birds are present."
The bulk of Kercher’s targeted BIRDWATCH samples have been from dabbling ducks like mallards and blue-winged teal, diving ducks like lesser scaup, and larger waterfowl like Canada, Ross’s, and snow geese. But news of two wild whooping crane carcasses testing positive for HPAI in Saskatchewan, Canada in Oct. of 2025 kick-started a pilot project for the BIRDWATCH lab.
Each fall, the approximately 550-strong flock of wild whooping cranes flies from their nesting grounds in Wood Buffalo National Park in northeastern Alberta, through the Great Plains, to Aransas National Wildlife Refuge on the Texas coast. Along the way, smaller groups of birds stopover at rivers, lakes, and crop fields to rest and refuel. Kercher and her research partners wanted to know if the endangered birds could come in contact with other waterbirds carrying the H5N1 virus or otherwise be exposed to the virus during their migration through the Central Flyway.
Because about a quarter of the wild whooping crane flock is tracked with radio telemetry devices, Kercher was able to plan an epic 10-day, 1,000-plus mile trip in Dec. 2025 that collected samples at 21 known whooping crane stopover sites in five states. Instead of physically taking samples from the endangered birds, she was able to follow the migration path and collect wetland sediment samples within feet of where a whooping crane had been months before.
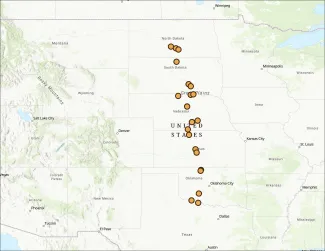
Kercher visited 21 known whooping crane stopover sites in five states to collect sediment samples for Highly Pathogenic Avian Influenza testing. “I looked at the whooping crane flyway and was impressed by the linear path. This is a bird that does not get distracted by latitude — it just goes straight down to the coast.”
“The virus may no longer be active, but its genetic material is bound up in the sediment,” Kercher said.
Five sediment samples were collected at each of the selected whooping crane sites, and each site’s samples were taken about 30 – 40 meters apart.
“I’m grateful for the rapid response and help from everyone in the flyway,” Kercher said. “Especially to those who braved the cold, muck, and wind to help me chip through the ice and navigate the mudflats.”

Lisa Kercher hands off a Red River sediment sample to Paxton Smith, a Wildlife Department biologist, as part of her 10-day visit to the Central Flyway. Kercher wore a glove on the hand she used to collect the sample to reduce the likelihood of cross-contamination between sites.
Along with the small containers of mud, muck, dirt, or sand scooped at each site, information about the wetland, its foot traffic, and sediment collection depth were recorded. Because Kercher’s whirlwind trip down the flyway lasted only 10 days, the 210 collected sediment samples were stored in the BIRDWATCH lab and later analyzed at St. Jude in Memphis, Tennessee.
There, the research team extracted RNA and ran preliminary PCR tests to detect the presence or absence of influenza genes, including Highly Pathogenic Avian Influenza.
“We got several hits across the flyway,” Kercher said.
Avian influenza wasn’t detected at every site but was detected in each of the five states, and the most current strain of Highly Pathogenic Avian Influenza, H5N1, was detected in three states, including Oklahoma. Sick birds, mostly geese, were observed in four of the five states.
“No one has done sediment testing for HPAI at the flyway scale,” Kercher said. “I was lucky to sample what I could — and it really begs the question of how this virus is moving since so many geese were sick, but rafts of thousands of healthy birds were also observed.
“By tracing the movement and evolution of HPAI across wild bird populations, the work by the team at St. Jude underscores how understanding diseases in nature can directly inform strategies to anticipate, prevent, and ultimately protect the world’s most vulnerable patients — children.”
To wrap up this pilot BIRDWATCH project within the Central Flyway, Kercher plans to visit the whooping crane’s wintering area along the Texas coast for additional sediment testing.

